Hello coders!! In this article, we will be learning about the Viterbi algorithm and its implementation in python. It is a dynamic programming algorithm used to find the most likely sequence of hidden states.
Key features of Viterbi Algorithm
- Dynamic programming algorithm
- Implemented to find the most likely sequence of hidden states (the Viterbi path)
- It reduces the number of computations by storing the calculations that are repeated
Mathematical Definition of Viterbi Algorithm:
A path X = (x1,x2,……xT) is generated which basically is a sequence of states x ∈ S = {s1,s2,……sK}. This generates the observation Y = (y1, y2, …., yT) with y ∈ O = {o1,o2,….oN}. Here, N is the possible number of observations in the observation space O.
In this we construct two 2D tables of size KxT
- Every element of table T1, i.e. T1[i,j] stores the probability of the most likely path so far X̂ = (x̂1, x̂2 …… x̂j) where x̂j = si will generate Y = (y1,y2 …yj)
- Every element of table T2, i.e. T2[i,j] stores x̂j-1 of the most likely path so far X̂ = (x̂1, x̂2 …… x̂j) ∀ j, 2 <= j <= T
- The elements of table T1[i,j] and T2[i,j] are filled in increasing order of K.j+i
- T1[i,j] = max(T1[k,j-1].Aki.Biyj)
- T2[i,j] = argmax(T1[k,j-1].Aki.Biyj)
Input:
- Observation space O = {o1,o2,…oN}
- State space S = {s1,s2,….,sk}
- An array consisting of initial probabilities π = (π1,π2….πk) where πi stores the probability x1 = si
- Sequence of observations Y = (y1, y2, …., yT)
- Transition matrix A of size K x K where A[i,j] stores the transition probability of transiting from state Si to Sj
- Emission matrix B of size K x N where B[i,j] stores the probability of observing Oj from state Si
Output:
The most likely hidden state sequence X=(x1,x2…xj)
Implementation of Viterbi Algorithm in Python with an example:
To illustrate the working of Viterbi algorithm we will be considering an example for better understanding.
Consider a small town where people are either:
- healthy
- have fever
Only the doctor can differentiate or identify the people having fever from healthy people. He does so by inquiring about their symptoms. People can have one of the following answers:
- normal
- dizzy
- cold
The doctor believes that the health condition of his patients operates as a discrete Markov chain.
The two states are:
- healthy
- fever
However, these states are ‘hidden’ from the doctor. The observations can be:
- normal
- dizzy
- cold
Thus, this forms a hidden Markov model which can be represented has:
observations = ("normal", "cold", "dizzy")
states = ("Healthy", "Fever")
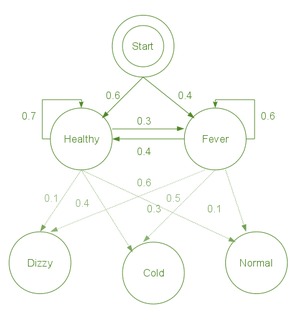
start_p = {"Healthy": 0.6, "Fever": 0.4}
trans_p = {
"Healthy": {"Healthy": 0.7, "Fever": 0.3},
"Fever": {"Healthy": 0.4, "Fever": 0.6},
}
emit_p = {
"Healthy": {"normal": 0.5, "cold": 0.4, "dizzy": 0.1},
"Fever": {"normal": 0.1, "cold": 0.3, "dizzy": 0.6},
}
- start_p: doctor’s analysis of patients’ state when he first pays a visit.
- trans_p: change of state
- In this example, we have considered a 30% change
- emission_p: how likely a state is determined based on observations
Now, consider a situation where a patient visits three days consecutively. All three days, he shows different symptoms (normal/cold/dizzy). The doctor’s question here will be: What is the most likely sequence of health conditions of the patient that would explain these observations?
This question is answered by Viterbi algorithm.
def viterbi_algorithm(observations, states, start_p, trans_p, emit_p):
V = [{}]
for st in states:
V[0][st] = {"prob": start_p[st] * emit_p[st][observations[0]], "prev": None}
for t in range(1, len(observations)):
V.append({})
for st in states:
max_tr_prob = V[t - 1][states[0]]["prob"] * trans_p[states[0]][st]
prev_st_selected = states[0]
for prev_st in states[1:]:
tr_prob = V[t - 1][prev_st]["prob"] * trans_p[prev_st][st]
if tr_prob > max_tr_prob:
max_tr_prob = tr_prob
prev_st_selected = prev_st
max_prob = max_tr_prob * emit_p[st][observations[t]]
V[t][st] = {"prob": max_prob, "prev": prev_st_selected}
for line in dptable(V):
print(line)
opt = []
max_prob = 0.0
best_st = None
for st, data in V[-1].items():
if data["prob"] > max_prob:
max_prob = data["prob"]
best_st = st
opt.append(best_st)
previous = best_st
for t in range(len(V) - 2, -1, -1):
opt.insert(0, V[t + 1][previous]["prev"])
previous = V[t + 1][previous]["prev"]
print ("The steps of states are " + " ".join(opt) + " with highest probability of %s" % max_prob)
def dptable(V):
yield " ".join(("%12d" % i) for i in range(len(V)))
for state in V[0]:
yield "%.7s: " % state + " ".join("%.7s" % ("%f" % v[state]["prob"]) for v in V)
The time complexity for the Viterbi Algorithm is O(N**2 * T) [n square times T]. Where N is the number of states and T is the sequence length.
Output:

Applications of Viterbi Algorithm:
- Decoding the convolutional codes used in both CDMA and GSM digital cellular
- Deep space communication
- Speech recognition and speech synthesis
- Keyword spotting
- Computational linguistics
Must Read
- Bitonic sort: Algorithm and Implementation in Python
- Fibonacci series in Python and Fibonacci Number Program
- Understanding Python Bubble Sort with examples
- Python Nmap Module Fully Explained with Programs
- Python is Not Recognized as an Internal or External Command
Conclusion:
In this article, we learned about the Viterbi Algorithm. We saw its implementation in Python, illustrated with the help of an example, and finally, we saw the various applications of the Viterbi Algorithm in modern technology.
However, if you have any doubts or questions, do let me know in the comment section below. I will try to help you as soon as possible.
Happy Pythoning!

Thanks for your detailed explanation helped me very. Can you please mention what is the time complexity of Viterbi alg?
The time complexity for the Viterbi Algorithm is O(N**2 * T) [n square times T]. Where N is the number of states and T is the sequence length.